Group17 proposal
Contents
Overview
Middle East Respiratory Syndrome Coronavirus (MERS-CoV) emerged as a global health concern in 2012 when the first human case was documented in Saudi Arabia. Then listed as one of the WHO Research and Development Blueprint priority pathogens, cases were reported in 27 countries across four continents. Imported cases into non-endemic countries such as France, Great Britain, the United States, and South Korea had caused secondary cases, thus highlighting the spread of MERS-CoV far beyond the countries where index cases originated. Reports in animals showed that viral circulation was far more widespread than suggested by human cases alone. In this project, we aim to analyse the spread of MERS-CoV and the factors affecting it's intensity.
Project Motivation
With the recent emergence of Covid virus, containing the epidemic requires an understanding of how corona virus spreads, and factors impacting the intensity of cases within and across regions. This project aims at delivering an R shiny app that first provides a basic understanding of the nature of the virus, e.g. the types of pathogens identified in MERS, the kinds of organisms which are susceptible to MERS contraction, and time series analysis to visualize the evolution of MERS outbreak across a 6-years time period (2012-2018). The detailed description of these variables are shown in the Section:Data Description. Geospatio-temporal analysis will be performed to identify the intensity of outbreak in different regions across time. Finally, we will further deep-dive into how certain factors intensifies the spread of the disease using spatial-join analysis.
Proposed Analytical Methods & Visualisation
1. Exploratory Data Analysis
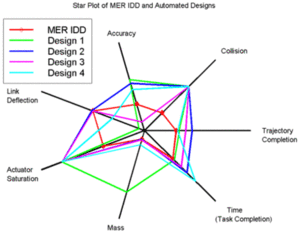
Radar Chart : A radar chart will be used to do multi-variate analysis, for instance, to show the different event types of the armed conflict over different countries in South Asia.
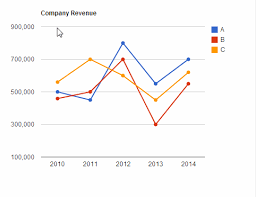
Line Chart : A line chart will be used to visualize the total number of incidents and the number of fatalities in those incidents over different periods of time.
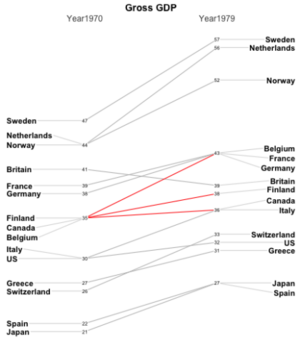
Slope Chart : Compares the ranking of countries over time and intensity of armed conflicts in South Asia over time to get a glimpse of the time series data.
2. Spatio-Temporal Analysis
Spatial Point (Describe point pattern analysis here) Kernel Density Plot 3D Chart (latitude, longitude and time)
3. Spatial Join Analysis Visualisations TBC
Project Timeline
Data Description
This database is publicly available online. Each of the 861 rows represents a unique occurrence of MERS-CoV. Rows containing an index, unspecified, or imported case represent a single case of MERS-CoV. Rows containing mammal and secondary cases may represent more than one case but are still unique geospatial occurrences.
| Data Fields | Description | Example | Datatype |
|---|---|---|---|
| nid | A unique identifier assigned to each publication that was extracted | 364253 | Numeric |
| occ_id | Unique identifier assigned to each occurrence of MERS-CoV. A single pdf may represent more than one occurrence. Each row will have its own occ_id, starting at 1 and numbered consecutively to 883. | 1 | Numeric |
| organism_type | What type of organism tested positive for MERS-CoV (human, mammal, or environmental). | human | Categorical |
| organism_specific | Specifies the exact organism that tested positive for MERS-CoV. Names are made consistent with Wilson and Reeder (2005) Mammal Species of the World. | Homo sapiens | Categorical |
| lat | This field records the latitude in decimal degrees. | 30.209423 | Numeric |
| long | This field records the longitude in decimal degrees. | 67.018009 | Numeric |
| pathogen | Name the pathogen identified (e.g. MERS-CoV, Bat Coronaviruses, and other MERS-CoV-like pathogens). | MERS-CoV | Categorical |
| patient_type | index, unspecified, NA, secondary, import, or absent. | index | Categorical |
| transmission_route | zoonotic, direct, unspecified, or animal-to-animal. | direct | Categorical |
| country | ISO3 code for country in which the case occurred. | KOR | Categorical |
| origin | Open-ended field to provide more details on the specific in-country location of MERS-CoV case. | Jordan | Categorical |
| loc_confidence | States the level of confidence that researchers had when assigning a geographic location to the MERS-CoV case (good or bad). | good | Categorical |
| month_start | Month that the occurrence(s) began. | 1 | Date Time |
| month_end | Month that the occurrence(s) ended. | 1 | Date Time |
| year_start | year that the occurrence(s) began. | 2013 | Year |
| year_end | year that the occurrence(s) ended. | 2012 | Year |
| year_accuracy | If years were reported, this field was assigned a value of ‘0’. If assumptions were required, this field was assigned a value of ‘1’. | 0 | Numeric |
Software Tools
- RStudio: https://rstudio.com/
Proposed R Packages
| Packages | Purpose |
|---|---|
| plotly() | To help with creating visuals for exploratory analysis |
| ggplot2() | To create elegant data visualizations using grammar of graphics |
| trelliscope() | To create interactive trelliscope displays |
| tidyverse() | To do data manipulation and exploration with dplyr() etc |
| gganimate() | To create plots with animation |
| leaflet() | To create maps within the application |
| spatstat() | To analyse spatial data |
| ads() | To analyse geographical data for spatial point pattern analysis |
| GeoXB() | To create interactive spatial exploratory data analysis |
| Shiny() | To create interactive web application for the final product |
References
1. https://acleddata.com/#/dashboard
Team Members
- Oishee Bhattacharyya
- Jaideep Ballani
- Denise Adele Chua Hui Shan